Comprehensive genomic profiling of associated biomarkers
GALEASTM Tumor (RUO)
Comprehensive genomic profiling of cancer-associated biomarkers for research use.
GALEAS Tumor (RUO) is for research use only and not for use in diagnostic procedures outside of the UK. For diagnostic applications, please see our GALEAS Tumor (UKCA) product. GALEAS Tumor - Comprehensive Genomic Profiling | Nonacus
Expertly curated content for CGP delivered through a single workflow
GALEAS Tumor is an expertly curated NGS panel developed in parallel with bioinformatics pipelines for comprehensive profiling of solid tumors in research studies. It targets 519 carefully selected genes associated with cancer, with content informed by the UK NHS national genomic test directory, NCCN, FDA and ESMO guidelines.
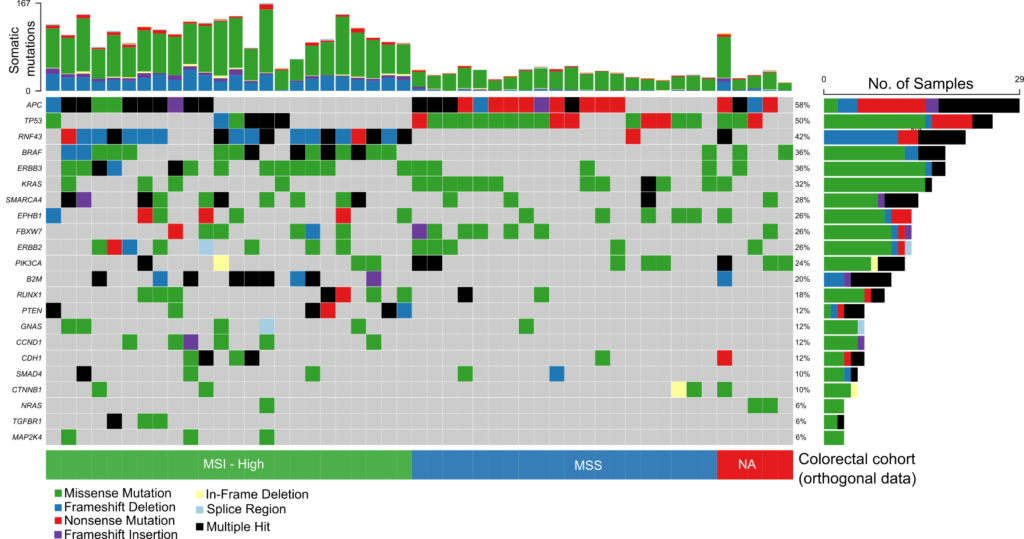
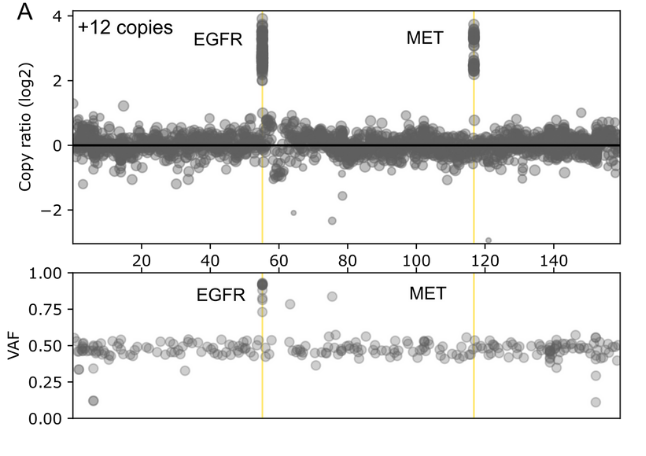
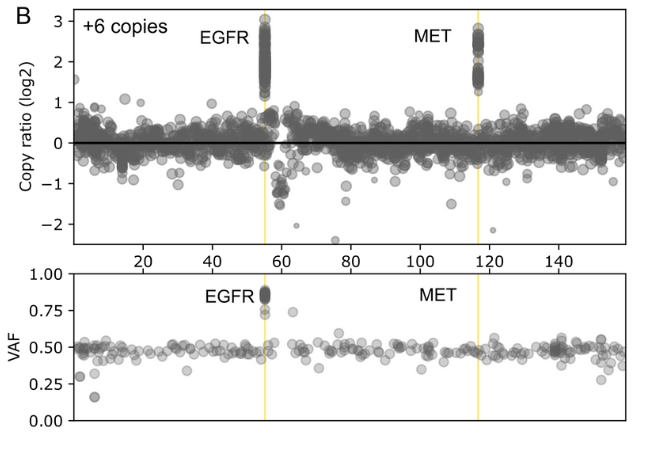
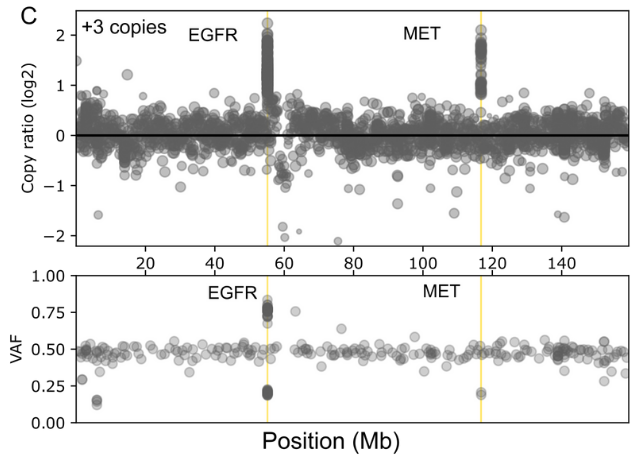
The comprehensive design covers genes found in both rare and common cancers and enables detection of a wide range of genetic variants including SNVs, INDELs, selected fusions and genome-wide CNVs in a single workflow
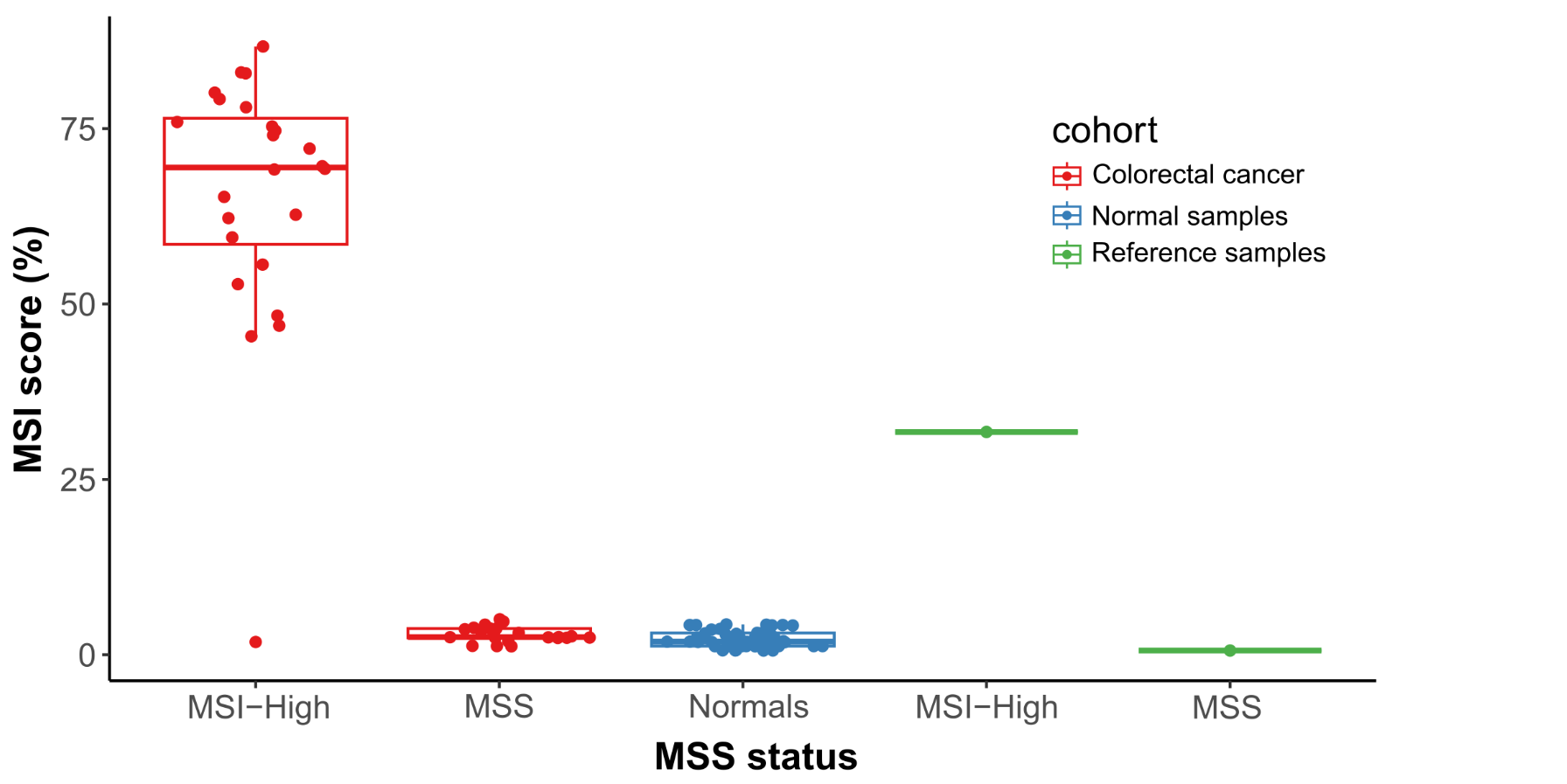
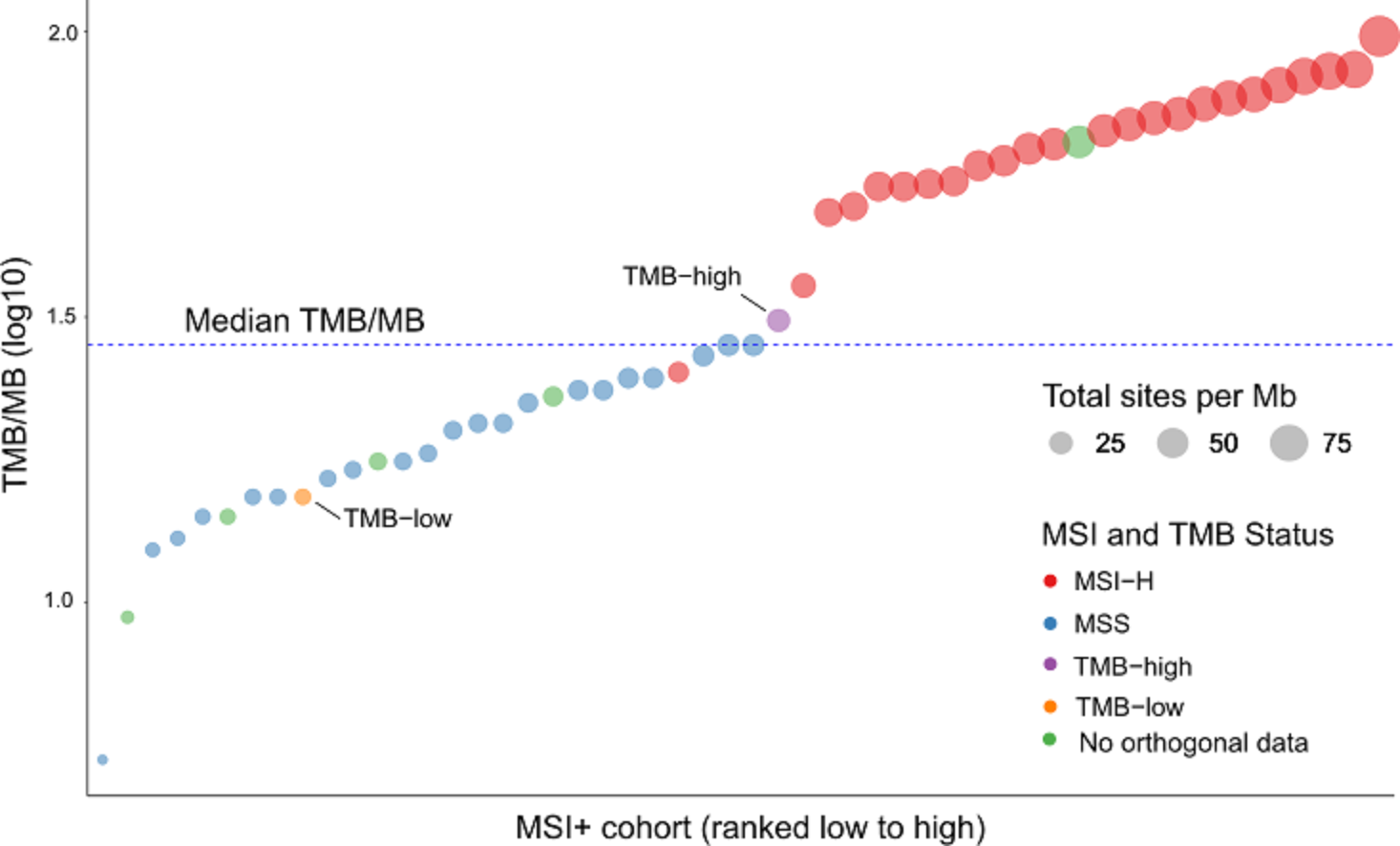
Probe enhancements support the assessment of other important biomarkers like Tumor Mutational Burden (TMB), Microsatellite Instability (MSI) and Homologous Recombination Deficiency (HRD).
Key Features
- Targets 519 genes for research into rare and common cancers including hereditary and pediatric cancers.
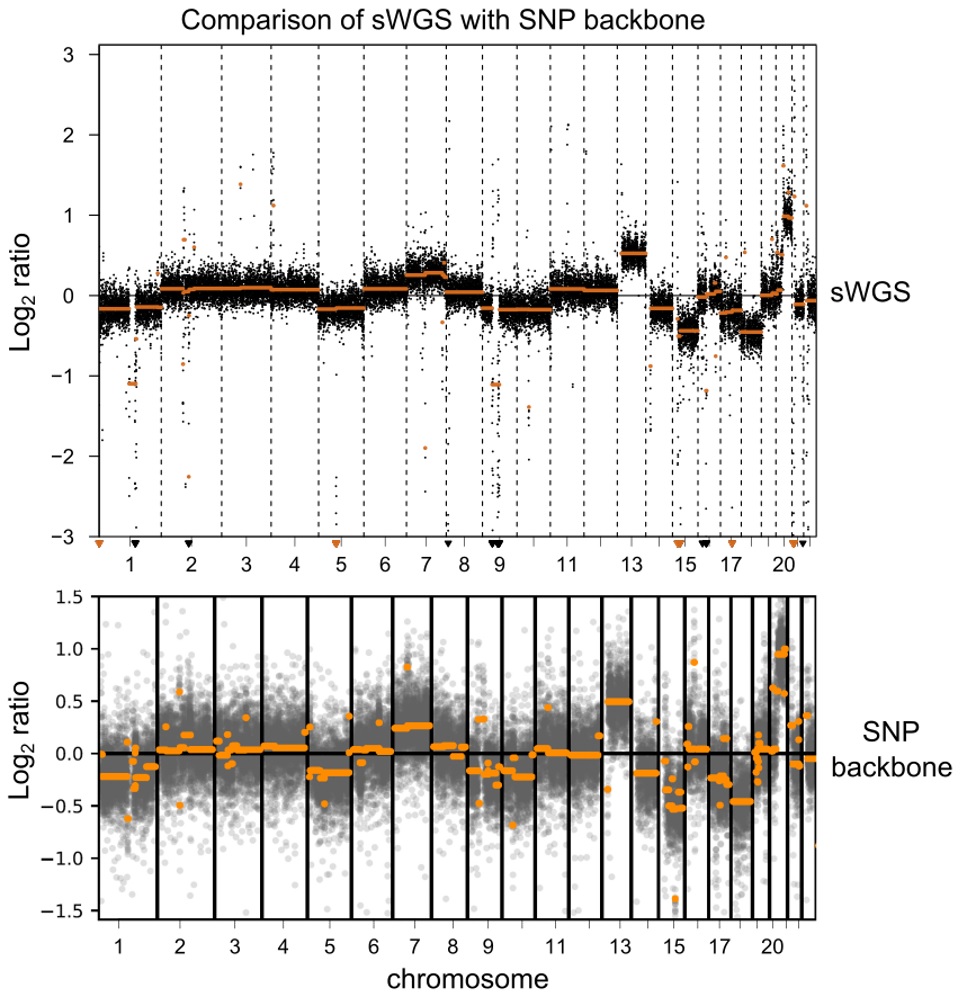
- Genome-wide SNP backbone delivers robust and reliable CNV calling.
- Probes enhancements allow MSI and TMB Immuno-oncology biomarker scoring.
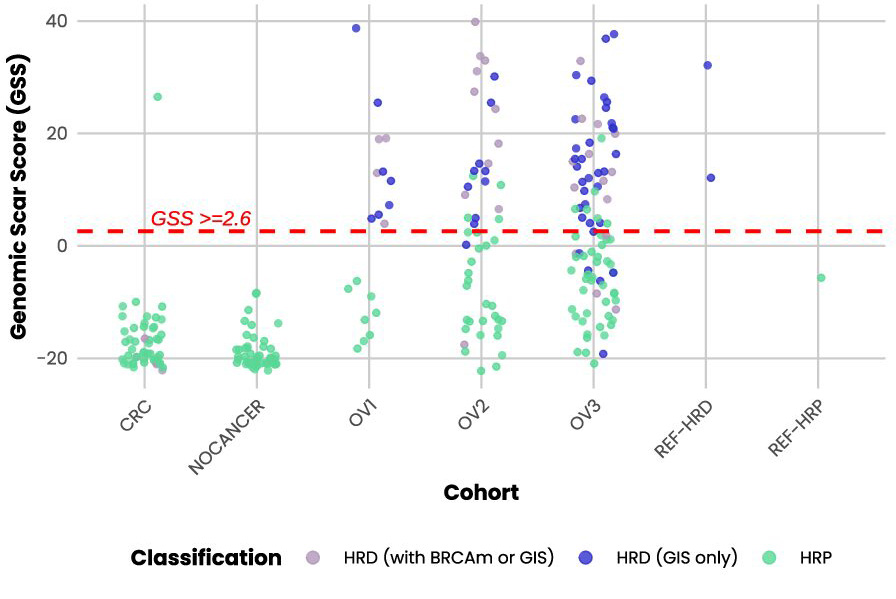
- BRCA variant calling and GI scoring deliver HRD assessment in a single workflow.
- Enhanced coverage of the 1p/19q co-deletion associated with Glioma supports brain cancer studies.
- Coverage of 10 Fusion/Structural rearrangements: ALK, BRAF, EGFR, FGFR2, FGFR3, NTRK1, NTRK2, RET, ROS1, TMPRSS2 reduces need for additional workflows.
- Sample identity tracking SNPs reduce potential errors in laboratory processes.
- Target 64 Pharmacogenomics (oncology) markers.
- Incorporates HLA design relevant for solid tumors.
Performance
Technical performance
High on-target rates and excellent uniformity of coverage deliver more efficient sequencing for comprehensive genomic profiling
GALEAS Tumor uses the Cell3 Target library preparation technology optimised by Nonacus to deliver a high percentage of on-target reads, low duplication rates and more uniform coverage.
This technical performance enables lower DNA inputs, less sequencing and enhanced recall and precision across more variants in research applications.
| Key quality indicator | GALEAS Tumor |
|---|---|
| Number of genes | 519 |
| Capture Panel Size (Mb) | 3.74 Mb |
| Gb required for mean 500x coverage (2x100bp PE) | 5 Gb |
| Percentage coverage ≥250x | 98% |
| Percentage on or near bait | 71% |
| Percent duplication | 9% |
| SNV recall | 100% |
| INDEL recall | 100% |
The GALEAS Tumor workflow is simple and easy

gDNA, FFPE DNA samples
Wide range of sample types including FFPE, frozen tissue and blood

Sample preparation
DNA extraction kits

Prepare libraries and enrich
GALEAS Tumor

Sequence
Illumina NGS Sequencing System

Call variants
GALEAS analysis software

Report
Generate reports for HRD, TMB and MSI

Interpret and report
Utilize third party tertiary software for interpretation and reporting
Why choose GALEAS Tumor?
Current and curated content
Understand the importance of SNVs, INDELs, selected fusions, CNVs, TMB, MSI and HRD in a wide range of cancers from common to rare in a single workflow.
Highly efficient capture chemistry
High on-target rates and excellent uniformity of coverage deliver more efficient sequencing and better sensitivity for comprehensive genomic profiling.
Supported by GALEAS software
Developed in parallel with the GALEAS Tumor panel, our GALEAS analysis software provides users with optimised bioinformatics for variant calling and supports integration into laboratories research workflows. The bioinformatics pipeline has not been validated for clinical diagnostic use.
Optimised library preparation
GALEAS Tumor library preparation is simple and easy compared to other targeted sequencing methods for hybridization and capture.
It requires as little as 10 ng of DNA and takes less than 10 hours, with less than two hours hands-on time. It is designed with multiple stop points to provide flexibility within laboratory processing. Library preparation can be run manually or automated (up to 96 samples in a single batch). Indexes are available for up to 384 samples to facilitate high throughput laboratories and to allow for flexible batch sizes.

Turnkey bioinformatics
Complimentary access to GALEAS cloud-based software delivers high quality variant calling and genome wide scores for research purposes
Cutting-edge bioinformatics pipelines were developed in tandem with, and specifically for GALEAS Tumor ensuring detection of all key variant types (SNV, INDEL, CNV and SV's) as well as genome wide scores for MSI and TMB and assessment of HRD. This turn-key software solution requires no specialist bioinformatic knowledge, enabling labs to integrate GALEAS Tumor into routine research use.
Including this optimised variant calling software with purchases of GALEAS Tumor provides researchers with tools to generate robust, reliable genomic information for research interpretation and analysis.
This software has not been validated for clinical diagnostic use.